Can a Person With Covid-19 Infect Other People Before Presenting Symptoms?
Abstract
We report temporal patterns of viral shedding in 94 patients with laboratory-confirmed COVID-nineteen and modeled COVID-19 infectiousness profiles from a separate sample of 77 infector–infectee manual pairs. Nosotros observed the highest viral load in throat swabs at the fourth dimension of symptom onset, and inferred that infectiousness peaked on or before symptom onset. We estimated that 44% (95% confidence interval, 30–57%) of secondary cases were infected during the index cases' presymptomatic stage, in settings with substantial household clustering, active case finding and quarantine outside the dwelling house. Disease control measures should be adjusted to account for probable substantial presymptomatic transmission.
Main
SARS-CoV-2, the causative agent of COVID-19, spreads efficiently, with a bones reproductive number of 2.two to 2.v determined in Wuhan1,2. The effectiveness of control measures depends on several primal epidemiological parameters (Fig. 1a), including the serial interval (duration betwixt symptom onsets of successive cases in a transmission concatenation) and the incubation period (fourth dimension between infection and onset of symptoms). Variation between individuals and transmission bondage is summarized by the incubation period distribution and the serial interval distribution, respectively. If the observed mean serial interval is shorter than the observed mean incubation flow, this indicates that a significant portion of manual may have occurred earlier infected persons accept adult symptoms. Pregnant presymptomatic transmission would probably reduce the effectiveness of control measures that are initiated past symptom onset, such as isolation, contact tracing and enhanced hygiene or employ of face masks for symptomatic persons.
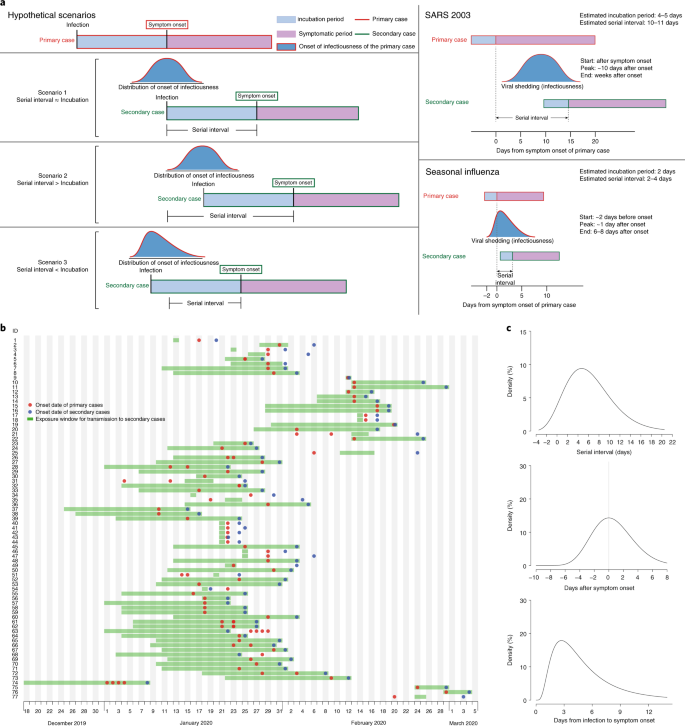
a, Schematic of the relation between different fourth dimension periods in the manual of infectious affliction. b, Human-to-homo transmission pairs of SAR-CoV-two virus (North = 77). We assumed a maximum exposure window of 21 days prior to symptom onset of the secondary cases. Detailed information on the transmission pairs and the source of information is summarized in Supplementary Tables two and three. c, Estimated serial interval distribution (top), inferred infectiousness profile (middle) and assumed incubation period (bottom) of COVID-19.
SARS (severe acute respiratory syndrome) was notable, because infectiousness increased around seven–10 days after symptom onset3,four. Onward transmission can be essentially reduced by containment measures such as isolation and quarantine (Fig. 1a)5. In contrast, influenza is characterized past increased infectiousness shortly around or fifty-fifty earlier symptom onsethalf dozen.
In this study, we compared clinical data on virus shedding with separate epidemiologic data on incubation periods and serial intervals betwixt cases in manual bondage, to describe inferences on infectiousness profiles.
Amidst 94 patients with laboratory-confirmed COVID-19 admitted to Guangzhou Eighth People's Infirmary, 47/94 (50%) were male, the median age was 47 years and 61/93 (66%) were moderately sick (with fever and/or respiratory symptoms and radiographic evidence of pneumonia), only none were classified as 'severe' or 'critical' on hospital admission (Supplementary Table 1).
A total of 414 throat swabs were collected from these 94 patients, from symptom onset up to 32 days subsequently onset. We detected high viral loads soon later symptom onset, which then gradually decreased towards the detection limit at about mean solar day 21. At that place was no obvious difference in viral loads across sex, age groups and disease severity (Fig. 2).
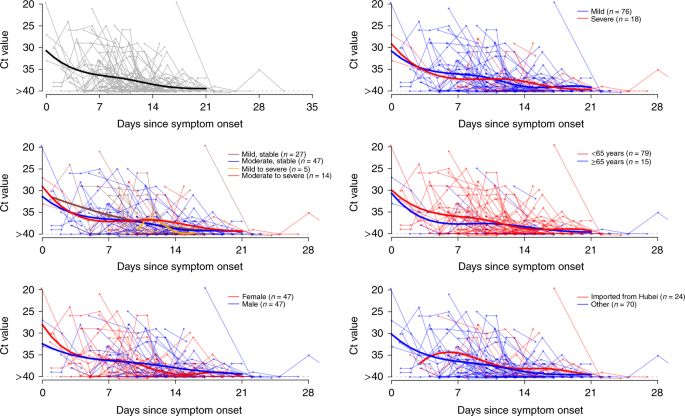
Viral load (threshold bike (Ct) values) detected by RT–PCR (PCR with reverse transcription) in throat swabs from patients infected with SARS-CoV-2 (Due north = 94), overall and stratified by disease severity, sex, age grouping and link to Hubei province. The detection limit was Ct = forty, which was used to indicate negative samples. The thick lines show the trend in viral load, using smoothing splines. We added some racket to the data points to avert overlaps.
Separately, based on 77 transmission pairs obtained from publicly available sources inside and outside mainland Communist china (Fig. 1b and Supplementary Table 2), the series interval was estimated to accept a mean of 5.8 days (95% confidence interval (CI), iv.viii–6.8 days) and a median of v.ii days (95% CI, four.1–half-dozen.four days) based on a fitted gamma distribution, with 7.vi% negative series intervals (Fig. 1c). Assuming an incubation period distribution of mean 5.2 days from a split up written report of early COVID-19 casesi, we inferred that infectiousness started from 12.three days (95% CI, v.9–17.0 days) earlier symptom onset and peaked at symptom onset (95% CI, –0.9–0.9 days) (Fig. 1c). Nosotros farther observed that only <0.ane% of transmission would occur earlier 7 days, ane% of transmission would occur before five days and 9% of manual would occur earlier 3 days prior to symptom onset. The estimated proportion of presymptomatic transmission (surface area under the curve) was 44% (95% CI, 30–57%). Infectiousness was estimated to decline chop-chop within 7 days. Viral load data were not used in the estimation but showed a similar monotonic decreasing pattern.
In sensitivity analysis, using the aforementioned estimating process but holding constant the start of infectiousness from 5, eight and 11 days earlier symptom onset, infectiousness was shown to peak at 2 days before to 1 solar day after symptom onset, and the proportion of presymptomatic transmission ranged from 37% to 48% (Extended Data Fig. i).
Finally, simulation showed that the proportion of short serial intervals (for example, <2 days) would be larger if infectiousness were assumed to beginning earlier symptom onset (Extended Data Fig. ii). Given the vii.6% negative serial intervals estimated from the infector–infectee paired data, start of infectiousness at to the lowest degree 2 days before onset and meridian infectiousness at 2 days before to 1 mean solar day after onset would be well-nigh consistent with this observed proportion (Extended Data Fig. 3).
Here, nosotros used detailed information on the timing of symptom onsets in transmission pairs to infer the infectiousness contour of COVID-19. We showed substantial transmission potential before symptom onset. Of note, most cases were isolated after symptom onset, preventing some post-symptomatic transmission. Even higher proportions of presymptomatic transmission of 48% and 62% accept been estimated for Singapore and Tianjin, where active example finding was implemented7. Places with active case finding would tend to have a higher proportion of presymptomatic transmission, mainly due to quick quarantine of shut contacts and isolation, thus reducing the probability of secondary spread afterwards on in the course of disease. In a chop-chop expanding epidemic wherein contact tracing/quarantine and perhaps even isolation are no longer feasible, or in locations where cases are not isolated outside the domicile, nosotros should therefore observe a lower proportion of presymptomatic transmission.
Our analysis suggests that viral shedding may begin v to 6 days earlier the advent of the starting time symptoms. Subsequently symptom onset, viral loads decreased monotonically, consistent with two recent studieseight,9. Another study from Wuhan reported that virus was detected for a median of twenty days (upward to 37 days among survivors) after symptom onset10, but infectiousness may pass up significantly 8 days afterwards symptom onset, as alive virus could no longer be cultured (according to Wölfel and colleagueseleven). Together, these results back up our findings that the infectiousness contour may more closely resemble that of influenza than of SARS (Fig. 1a), although we did not have data on viral shedding before symptom onsethalf dozen,12. Our results are as well supported by reports of asymptomatic and presymptomatic manualthirteen,14.
For a reproductive number of 2.5 (ref. two), contact tracing and isolation lone are less likely to be successful if more than xxx% of transmission occurred before symptom onset, unless >ninety% of the contacts tin be traced15. This is more likely achievable if the definition of contacts covers 2 to iii days prior to symptom onset of the alphabetize case, as has been done in Hong Kong and mainland China since belatedly February. Even when the control strategy is shifting away from containment to mitigation, contact tracing would notwithstanding exist an of import measure, such equally when there are super-spreading events that may occur in high-gamble settings including nursing homes or hospitals. With a substantial proportion of presymptomatic transmission, measures such as enhanced personal hygiene and social distancing for all would likely be the fundamental instruments for community disease control.
Our study has several limitations. Outset, symptom onset relies on patient call up afterward confirmation of COVID-xix. The potential call back bias would probably have tended toward the direction of nether-ascertainment, that is, delay in recognizing get-go symptoms. As long as these biases did not differ systematically between infector and infectee, the serial interval estimate would not exist essentially afflicted. Nevertheless, the incubation period would have been overestimated, and thus the proportion of presymptomatic transmission artifactually inflated. Second, shorter serial intervals than those reported here take been reported, only such estimates lengthened when restricted to infector–infectee pairs with more sure transmission links16. Finally, the viral shedding dynamics were based on information for patients who received handling co-ordinate to nationally promulgated protocols, including combinations of antivirals, antibiotics, corticosteroids, immunomodulatory agents and Chinese medicine preparations, which could take modified the shedding dynamical patterns.
In decision, we have estimated that viral shedding of patients with laboratory-confirmed COVID-19 peaked on or before symptom onset, and a substantial proportion of transmission probably occurred before first symptoms in the index case. More inclusive criteria for contact tracing to capture potential manual events 2 to iii days before symptom onset should be urgently considered for effective control of the outbreak.
Methods
Sources of information
Guangzhou Eighth People's Infirmary in Guangdong, China was designated as one of the specialized hospitals for treating patients with COVID-19 at both city and provincial levels on twenty Jan 2020. Later on that, many people with COVID-19 were admitted via fever clinics, the hospital emergency room or afterwards confirmation of cases from community epidemiological investigation carried out by the Guangzhou Center for Disease Control and Prevention, or transferred from other hospitals. The commencement confirmed patient with COVID-19 was admitted on 21 January 2020, just in the initial phase, patients suspected to have COVID-19 were also admitted. Nosotros identified all suspected and confirmed COVID-19 cases admitted from 21 January 2022 to fourteen February 2022 and collected throat swabs in each instance. Patients included those who traveled from Wuhan or Hubei to Guangzhou too as locals, with cases ranging from asymptomatic, mild to moderate at access.
The samples were tested past N-gene-specific quantitative RT–PCR assay as previously described17. To empathize the temporal dynamics of viral shedding and exclude non-confirmed COVID-19 cases, nosotros selected 94 patients who had at least one positive result (cycle threshold (Ct) value < twoscore) in their throat samples. Series samples were collected from some only not all patients for clinical monitoring purposes.
We nerveless information reported on possible human-to-human transmission pairs of patients with laboratory-confirmed COVID-xix from publicly available sources, including announcements made by government wellness agencies and media reports in mainland Mainland china and countries/regions outside China. A transmission pair was defined as two confirmed COVID-19 cases identified in the epidemiologic investigation by showing a clear epidemiologic link with each other, such that one case (infectee) was highly likely to take been infected by the other (infector), by fulfilling the following criteria: (1) the infectee did not written report a travel history to an surface area affected past COVID-19 or any contact with other confirmed or suspected COVID-xix cases except for the infector within 14 days before symptom onset; (2) the infector and infectee were not identified in a patient cluster where other COVID-19 cases had besides been confirmed; and (3) the infector and infectee pair did not share a common source of exposure to a COVID-19 case or a place where at that place were COVID-19 case(due south) reported. We excluded possible transmission pairs without a clear exposure history reported prior to symptom onset. Information of possible transmission pairs of COVID-19 were extracted, including age, sex, location, date of symptom onset, type or relationship between the pair cases and time of contact of the cases.
Statistical analysis
We analyzed two divide data sets—clinical and epidemiologic—to assess presymptomatic infectiousness. First, we assessed longitudinal viral shedding data from patients with laboratory-confirmed COVID-19 starting from symptom onset, where viral shedding during the first few days after illness onset could exist compared with the inferred infectiousness. 2d, the serial intervals from clear transmission chains, combined with data on the incubation period distribution, were used to infer the infectiousness contour, as described in the following.
We present SARS-CoV-2 viral loads in the throat swabs of each patient by day of symptom onset. To aid visualization, a smoothing spline was fitted to the Ct values to summarize the overall trend. Specifically, a generalized additive model, E(Y) =β 0 +s(t), with an identity link was fitted, where Y are the Ct values, β 0 is the intercept and s(t) is a cubic spline evaluated at t days after symptom onset. We also compared the viral load by disease severity, age, sexual practice and travel history from Hubei.
We fitted a gamma distribution to the transmission pairs data to approximate the serial interval distribution. We used a published estimate of the incubation period distribution to infer infectiousness with respect to symptom onset from the beginning 425 patients with COVID-xix in Wuhan with detailed exposure history1. We considered that infected cases would become infectious at a sure time signal before or after illness onset (t S1). Infectiousness—that is, transmission probability to a secondary case—would then increase until reaching its summit (Fig. 1). The manual consequence would occur at fourth dimension t I with a probability described by the infectiousness profile β c (t I −t S1) relative to the illness onset date, assuming a gamma distribution β(t) with a time shift c to permit for first of infectiousness c days prior to symptom onset; that is, β c (t) =β(t +c). The secondary case would and then show symptoms at fourth dimension t S2, later the incubation period that is causeless to follow a lognormal distribution g(t S2 −t I). Hence the observed series intervals distribution f(t S2 −t S1) would exist the convolution between the infectiousness profile and incubation period distribution. We constructed a likelihood function based on the convolution, which was fitted to the observed serial intervals, allowing for the commencement of infectiousness around symptom onset and window of symptom onset (t S1l, t S1u), given past
$$50\left( {t_{\rm{S1u}},t_{\rm{S1l}},t_{\rm{S2}}|\theta } \correct) = \mathop {\int}\limits_{t_{\rm{S1l}}}^{t_{\rm{S1u}}} {\mathop {\int}\limits_{ - \infty }^{t_{\rm{S}ii}} {\beta _c} } \left( {t_{\rm{I}} - t_{\rm{S}1}} \right)g(t_{\rm{South}2} - t_{\rm{I}}){\rm{d}}t_{\rm{I}}{\rm{d}}t_{\rm{Due south}ane}$$
A normalization factor tin be added to account for the uncertainty in the symptom-onset dates of the index cases. Assuming a uniform distribution, the likelihood would differ only by a multiplicative constant and give the same estimates.
Parameters θ, including the gamma distribution parameters and the start of infectiousness, were estimated using maximum likelihood. The 95% CIs were obtained by bootstrapping with 1,000 replications. We also performed sensitivity analyses by fixing the start of infectiousness from days 5, 8 and 11 before symptom onset and inferred the infectiousness contour.
As an additional check, nosotros simulated the expected series intervals bold the aforementioned aforementioned incubation period but two different infectiousness profiles, where infectiousness started on the same mean solar day and from 2 days before symptom onset, respectively. A recent study isolated live infectious SARS-CoV-2 virus from patients with COVID-19 up to eight days later on symptom onset11, thus nosotros assumed the same duration of infectiousness. We also assumed that infectiousness peaked on the mean solar day of symptom onset. The timing of transmission to secondary cases was faux co-ordinate to the infectiousness profile using a lognormal and exponential distribution, respectively, where the serial intervals were estimated as the sum of the onset to transmission interval and the incubation period. We drew random samples for the transmission time relative to symptom onset of the infector T I ≈β c (t), and also the incubation period T inc ≈f(t), then the simulated serial interval was T I +T inc. Nosotros also performed simulation considering combinations of unlike infectiousness profiles, with start of infectiousness vii days earlier to 3 days after symptom onset, and pinnacle infectiousness as well 7 days before to iii days afterwards symptom onset. We nowadays the distribution of the serial intervals and proportion of negative serial intervals over 10,000 simulations.
All statistical analyses were conducted in R version 3.half dozen.3 (R Development Core Squad).
Ideals approval
Data collection and analysis were required past the National Health Commission of the People'southward Democracy of China to be part of a continuing public health outbreak investigation.
Reporting Summary
Farther information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Detailed transmission pairs data in this study are provided in the Supplementary Information and viral shedding data volition exist available upon request and approval by a data access committee. The information access commission comprises leadership of the Guangzhou Eighth People'due south Infirmary and the Guangzhou Health Commission. At that place is no restriction to information access.
Lawmaking availability
We provided the code for generating Fig. 1c in the Supplementary Information and at https://github.com/ehylau/COVID-19. Other codes are bachelor upon request to the respective author.
Alter history
-
07 August 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41591-020-1016-z
References
-
Li, Q. et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Engl. J. Med. 382, 1199–1207 (2020).
-
Wu, J. T., Leung, K. & Leung, G. M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study. Lancet 395, 689–697 (2020).
-
Peiris, J. S. et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet 361, 1767–1772 (2003).
-
Pitzer, V. Eastward., Leung, G. M. & Lipsitch, M. Estimating variability in the transmission of astringent astute respiratory syndrome to household contacts in Hong Kong, China. Am. J. Epidemiol. 166, 355–363 (2007).
-
Riley, S. et al. Transmission dynamics of the etiological amanuensis of SARS in Hong Kong: impact of public health interventions. Scientific discipline 300, 1961–1966 (2003).
-
Ip, D. K. et al. Viral shedding and transmission potential of asymptomatic and paucisymptomatic influenza virus infections in the customs. Clin. Infect. Dis. 64, 736–742 (2017).
-
Ganyani, T. et al. Estimating the generation interval for COVID-19 based on symptom onset information. Preprint at medRxiv https://doi.org/10.1101/2020.03.05.20031815 (2020).
-
Zou, L. et al. SARS-CoV-two viral load in upper respiratory specimens of infected patients. Due north. Engl. J. Med. 382, 1177–1179 (2020).
-
To, Thousand.K.-Westward. et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(20)30196-1 (2020).
-
Zhou, F. et al. Clinical course and hazard factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395, 1054–1062 (2020).
-
Wölfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature https://doi.org/10.1038/s41586-020-2196-x (2020).
-
Tsang, T. One thousand. et al. Influenza A virus shedding and infectivity in households. J. Infect. Dis. 212, 1420–1428 (2015).
-
Bai, Y. et al. Presumed asymptomatic carrier transmission of COVID-19. JAMA https://doi.org/x.1001/jama.2020.2565 (2020).
-
Tong, Z. D. et al. Potential presymptomatic manual of SARS-CoV-2, Zhejiang Province, People's republic of china, 2020. Emerg. Infect. Dis.https://doi.org/10.3201/eid2605.200198 (2020).
-
Hellewell, J. et al. Feasibility of controlling COVID-19 outbreaks past isolation of cases and contacts. Lancet Glob. Wellness 8, e488–e496 (2020).
-
Nishiura, H., Linton, North. M. & Akhmetzhanov, A. R. Serial interval of novel coronavirus (COVID-xix) infections. Int. J. Infect. Dis. 93, 284–286 (2020).
-
Chen, W. et al. Detectable 2019-nCoV viral RNA in blood is a stiff indicator for the farther clinical severity. Emerg. Microbes Infect. 9, 469–473 (2020).
Acknowledgements
This work was supported past Department of Scientific discipline and Technology of Guangdong Province (projection no. 2020B111108001, to F.L.) and a commissioned grant from the Health and Medical Research Fund from the Government of the Hong Kong Special Administrative Region (to B.J.C.).
Author information
Affiliations
Contributions
Ten. He, Due east.H.Y.50., P.W., B.J.C., F.L. and Thousand.Yard.L. conceived and designed the written report. 10. He, X.D., J.W., Y.G., X.T., X.M., Y.C. and B.50. were responsible for clinical intendance and collected all biomaterials. W.C. and F.H. carried out laboratory testing. Q.Z., One thousand.Z. and Y.Westward. collected and collated linked clinical–epidemiologic information. L.Z., F.Z. and F.L. supervised and coordinated all aspects of the study at Guangzhou 8th People'due south Hospital. P.W., Ten. Hao, Y.C.L. and J.Y.W. collected and verified all infector–infectee transmission data. East.H.Y.50., B.J.C. and Chiliad.Yard.50. wrote the start draft. All authors contributed to data estimation, critical revision of the manuscript and canonical the last version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Boosted information
Peer review information Alison Farrell and João Monteiro were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial squad.
Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. i Inferred infectiousness profile.
Infectiousness was assumed to start from 5 days (height) to eight days (middle) and 11 days (bottom) earlier symptom onset.
Extended Data Fig. 2 False serial intervals.
False series intervals assuming infectiousness started on the same day of symptom onset (top panel) and from two days earlier symptom onset (bottom panel) to about 10 days after symptom onset. Both scenarios assumed that infectiousness peaked on the first twenty-four hours of symptom onset.
Extended Data Fig. iii Faux proportions of negative serial intervals.
Simulated proportions of negative serial intervals assuming get-go of infectiousness and peak infectiousness from seven days earlier symptom onset to 3 days later symptom onset. From the estimated serial interval distribution based on infector-infectee pairs, vii.half-dozen% of the serial intervals were negative. Grey area represents the implausible range where the peak infectious is earlier than the start of infectiousness.
Supplementary information
Rights and permissions
About this article
Cite this article
He, 10., Lau, Eastward.H.Y., Wu, P. et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med 26, 672–675 (2020). https://doi.org/10.1038/s41591-020-0869-5
-
Received:
-
Accustomed:
-
Published:
-
Issue Date:
-
DOI : https://doi.org/x.1038/s41591-020-0869-five
Further reading
Source: https://www.nature.com/articles/s41591-020-0869-5
0 Response to "Can a Person With Covid-19 Infect Other People Before Presenting Symptoms?"
Publicar un comentario